Precomputed forward model results
We provide precomputed fluence and sensitivity files for the example datasets. These are created by this notebook and can be obtained through cedalion.data.get_precomputed_sensitivity.
The second part of the notebook visualizes the currently available sensitivities.
[1]:
# This cells setups the environment when executed in Google Colab.
try:
import google.colab
!curl -s https://raw.githubusercontent.com/ibs-lab/cedalion/dev/scripts/colab_setup.py -o colab_setup.py
# Select branch with --branch "branch name" (default is "dev")
%run colab_setup.py
except ImportError:
pass
[2]:
# set to true, to actually compute fluence and sensitivity files
RUN_COMPUTATION = False
[3]:
import pyvista as pv
pv.set_jupyter_backend('static')
from functools import lru_cache
import os
from pathlib import Path
import xarray as xr
import cedalion.data
import cedalion.dot as dot
from cedalion.io.forward_model import FluenceFile, load_Adot
from cedalion.vis.anatomy import sensitivity_matrix
xr.set_options(display_expand_data=False);
[4]:
def compute_fluence_mcx(geo3d_snapped_ijk, meas_list, head, output_file):
fwm = dot.ForwardModel(
head, geo3d_snapped_ijk, meas_list
)
fwm.compute_fluence_mcx(output_file)
def compute_sensitivity(geo3d_snapped_ijk, meas_list, head, fluence_fname, sensitivity_fname):
fwm = dot.ForwardModel(
head, geo3d_snapped_ijk, meas_list
)
fwm.compute_sensitivity(fluence_fname, sensitivity_fname)
def get_fnirs_dataset(dataset):
if dataset == "fingertappingDOT":
rec = cedalion.data.get_fingertappingDOT()
return rec.geo3d, rec._measurement_lists["amp"]
elif dataset == "fingertapping":
rec = cedalion.data.get_fingertapping()
return rec.geo3d, rec._measurement_lists["amp"]
elif dataset == "nn22_resting":
rec = cedalion.data.get_nn22_resting_state()
return rec.geo3d, rec._measurement_lists["amp"]
elif dataset == "ninja_cap_56x144":
geo3d, landmarks, meas_list = cedalion.data.get_ninja_cap_probe()
geo3d = xr.concat((geo3d, landmarks), dim="label")
geo3d = geo3d.pint.quantify("mm")
return geo3d, meas_list
elif dataset == "ninja_uhd_cap_164x496":
geo3d, landmarks, meas_list = cedalion.data.get_ninja_uhd_cap_probe()
geo3d = xr.concat((geo3d, landmarks), dim="label")
geo3d = geo3d.pint.quantify("mm")
return geo3d, meas_list
def snap(head, geo3d, dataset):
if dataset in ["fingertapping", "fingertappingDOT"]:
# right handed; too few fiducials
geo3d_snapped_ijk = head.align_and_snap_to_scalp(
geo3d, mode="trans_rot_isoscale"
)
elif dataset in ["nn22_resting", "ninja_cap_56x144", "ninja_uhd_cap_164x496"]:
# left handed
geo3d_snapped_ijk = head.align_and_snap_to_scalp(geo3d, mode="general")
return geo3d_snapped_ijk
def plot_sensitivity(dataset, headmodel, Adot=None):
sensitivity_fname = f"sensitivity_{dataset}_{headmodel}.nc"
if Adot is None:
Adot = load_Adot(sensitivity_fname)
head = cedalion.dot.get_standard_headmodel(headmodel)
geo3d, meas_list = get_fnirs_dataset(dataset)
geo3d_snapped_ijk = snap(head, geo3d, dataset)
plotter = sensitivity_matrix.Main(
sensitivity=Adot,
brain_surface=head.brain,
head_surface=head.scalp,
labeled_points=geo3d_snapped_ijk,
)
plotter.plot(high_th=0, low_th=-3)
plotter.plt.show()
def compute_fluence_and_sensitivity(dataset : str, headmodel : str):
geo3d, meas_list = get_fnirs_dataset(dataset)
head = cedalion.dot.get_standard_headmodel(headmodel)
geo3d_snapped_ijk = snap(head, geo3d, dataset)
fluence_fname = f"fluence_{dataset}_{headmodel}.h5"
sensitivity_fname = f"sensitivity_{dataset}_{headmodel}.nc"
if not Path(fluence_fname).exists():
print(f"computing fluence for {dataset} {headmodel}")
compute_fluence_mcx(geo3d_snapped_ijk, meas_list, head, fluence_fname)
else:
print(f"fluence for {dataset} {headmodel} exists")
if not Path(sensitivity_fname).exists():
print(f"computing sensitivity for {dataset} {headmodel}")
compute_sensitivity(geo3d_snapped_ijk, meas_list, head, fluence_fname, sensitivity_fname)
else:
print(f"sensitivity for {dataset} {headmodel} exists")
[5]:
if RUN_COMPUTATION:
for dataset in [
"fingertapping",
"fingertappingDOT",
"nn22_resting",
"ninja_cap_56x144",
"ninja_uhd_cap_164x496",
]:
for headmodel in ["colin27", "icbm152"]:
compute_fluence_and_sensitivity(dataset, headmodel)
plot_sensitivity(dataset, headmodel)
Visualize precomputed sensitivities
[6]:
for dataset in ["fingertapping", "fingertappingDOT", "nn22_resting", "ninja_cap_56x144", "ninja_uhd_cap_164x496"]:
for headmodel in ["colin27", "icbm152"]:
display(f"{dataset=} {headmodel=}")
Adot = cedalion.data.get_precomputed_sensitivity(dataset, headmodel)
plot_sensitivity(dataset, headmodel, Adot)
"dataset='fingertapping' headmodel='colin27'"
Downloading file 'sensitivity_fingertapping_colin27.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_fingertapping_colin27.nc' to '/home/runner/.cache/cedalion/dev'.
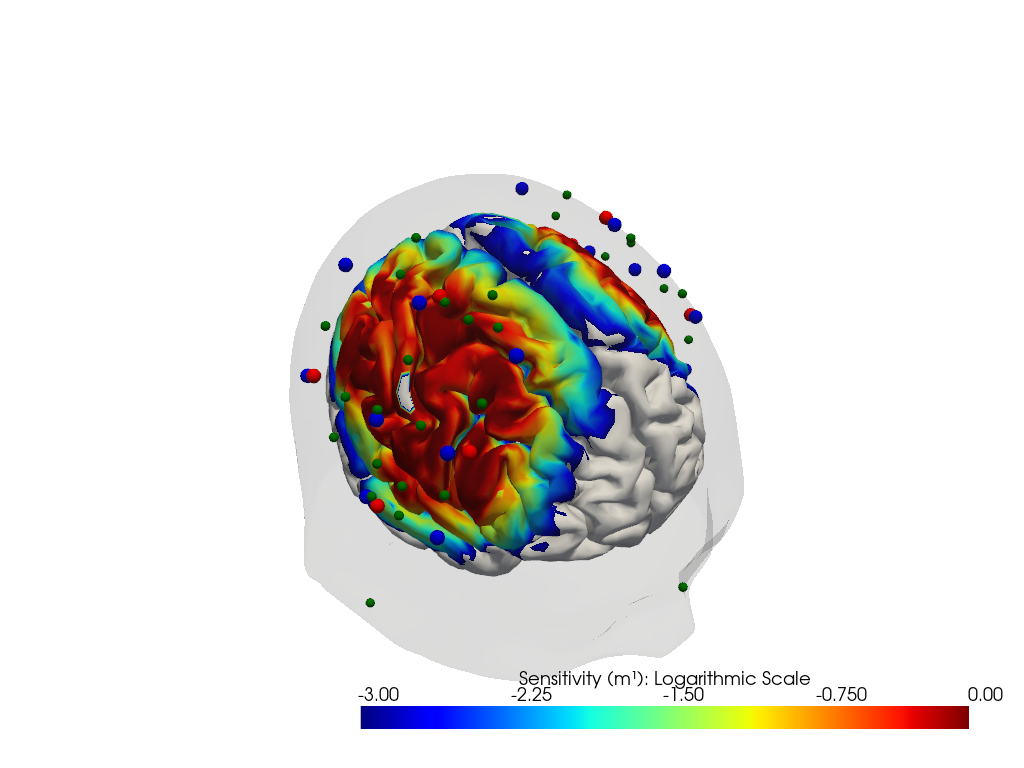
"dataset='fingertapping' headmodel='icbm152'"
Downloading file 'sensitivity_fingertapping_icbm152.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_fingertapping_icbm152.nc' to '/home/runner/.cache/cedalion/dev'.

"dataset='fingertappingDOT' headmodel='colin27'"
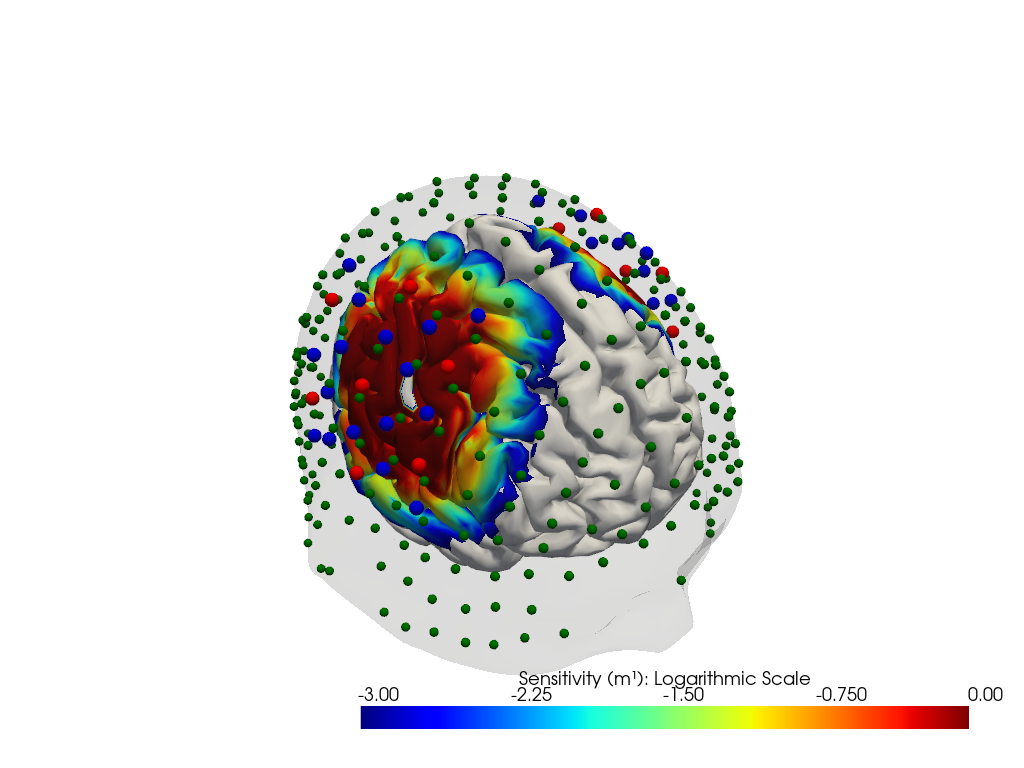
"dataset='fingertappingDOT' headmodel='icbm152'"
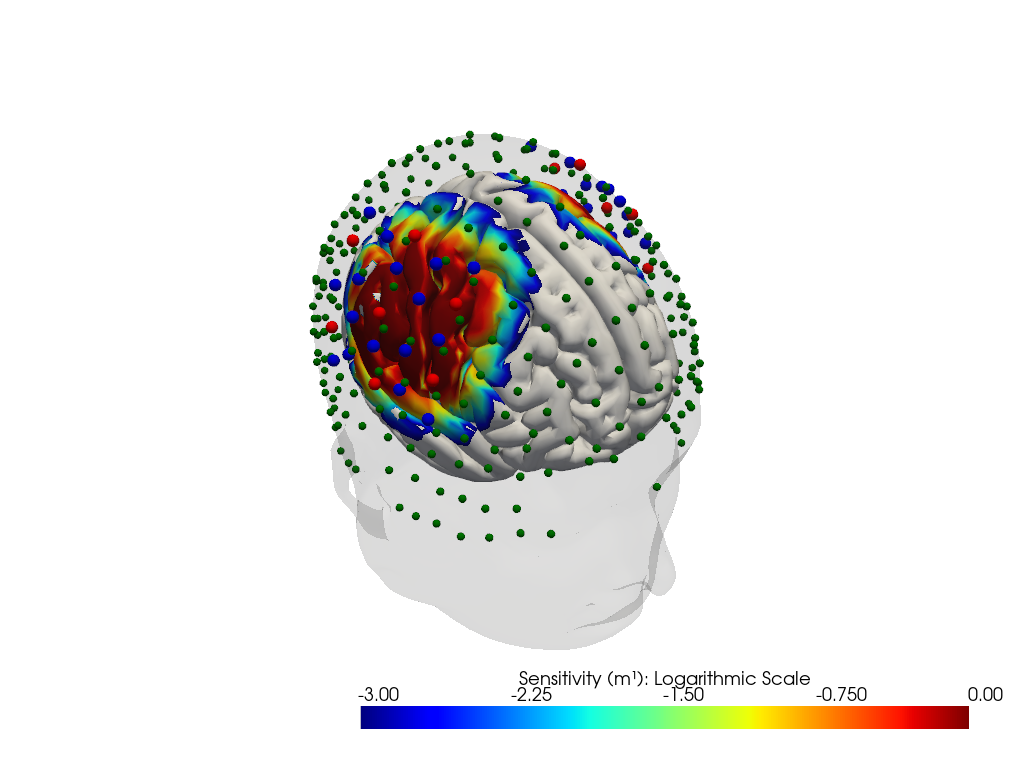
"dataset='nn22_resting' headmodel='colin27'"

"dataset='nn22_resting' headmodel='icbm152'"
Downloading file 'sensitivity_nn22_resting_icbm152.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_nn22_resting_icbm152.nc' to '/home/runner/.cache/cedalion/dev'.
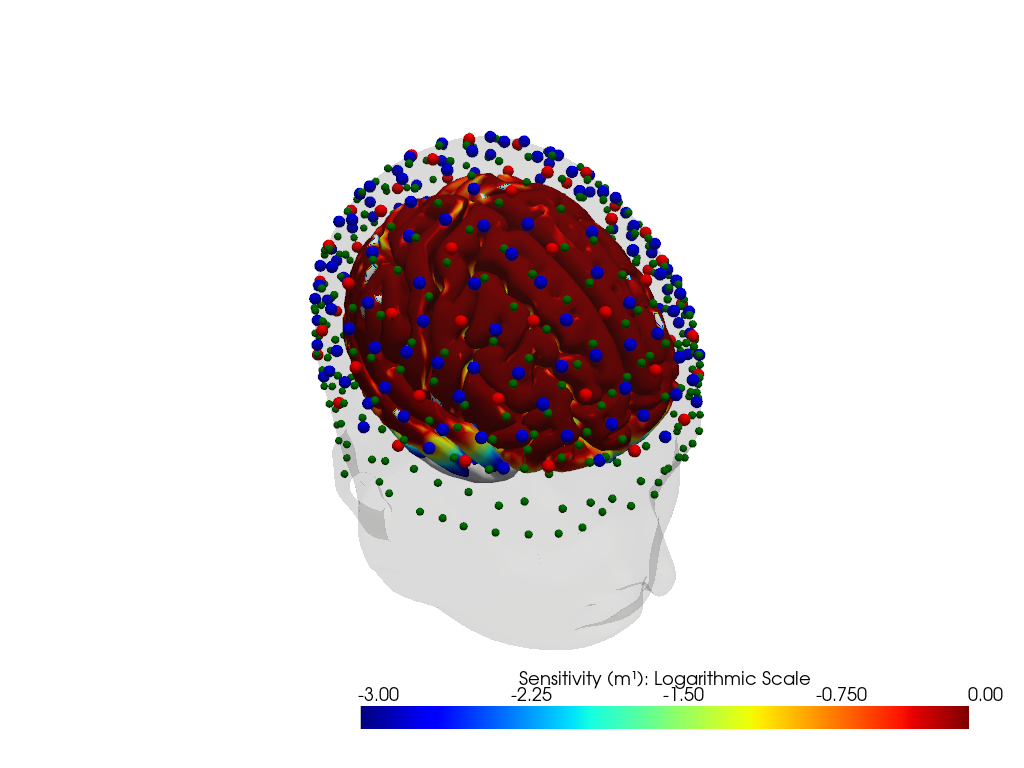
"dataset='ninja_cap_56x144' headmodel='colin27'"
Downloading file 'sensitivity_ninja_cap_56x144_colin27.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_ninja_cap_56x144_colin27.nc' to '/home/runner/.cache/cedalion/dev'.
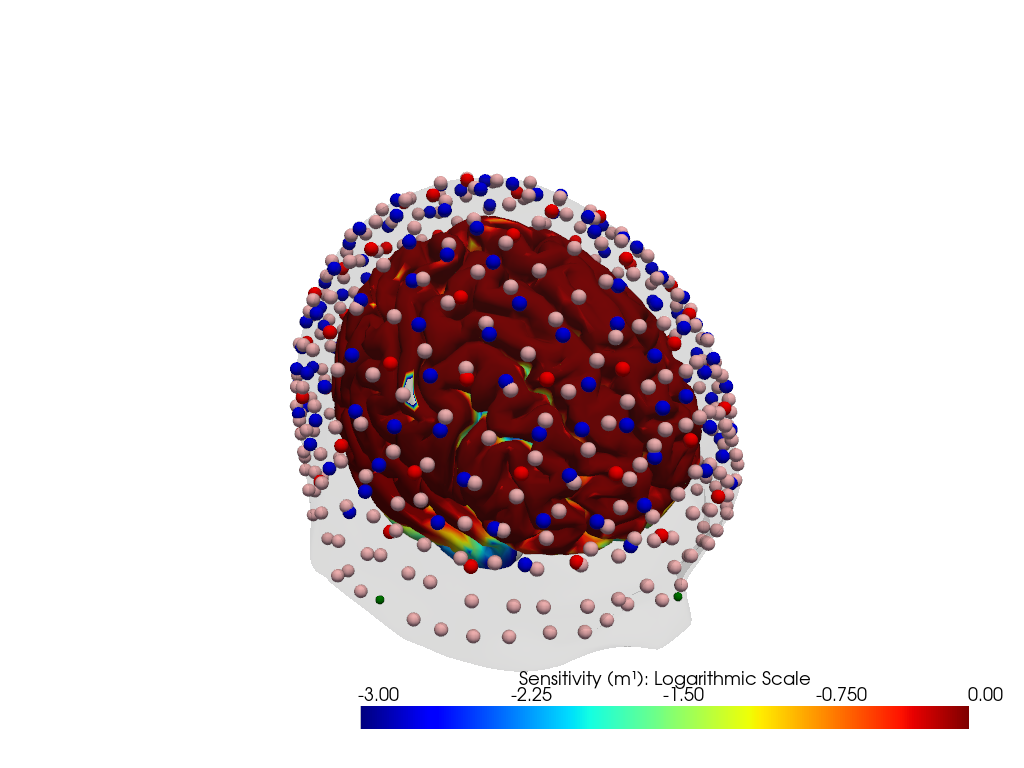
"dataset='ninja_cap_56x144' headmodel='icbm152'"
Downloading file 'sensitivity_ninja_cap_56x144_icbm152.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_ninja_cap_56x144_icbm152.nc' to '/home/runner/.cache/cedalion/dev'.

"dataset='ninja_uhd_cap_164x496' headmodel='colin27'"
Downloading file 'sensitivity_ninja_uhd_cap_164x496_colin27.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_ninja_uhd_cap_164x496_colin27.nc' to '/home/runner/.cache/cedalion/dev'.
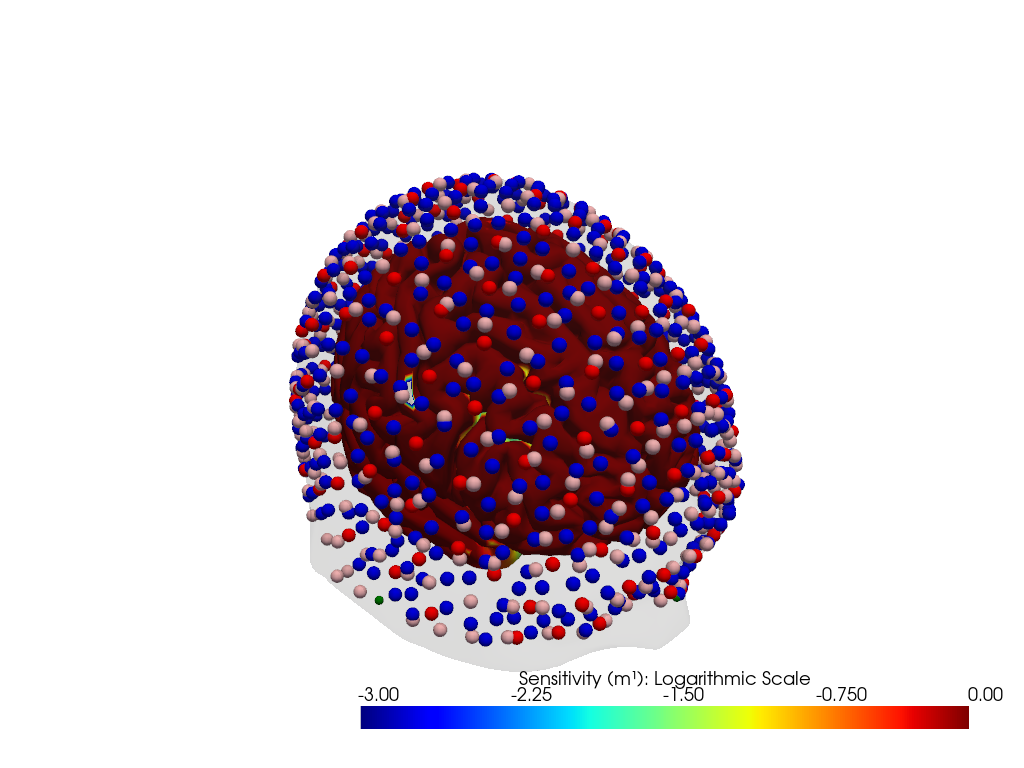
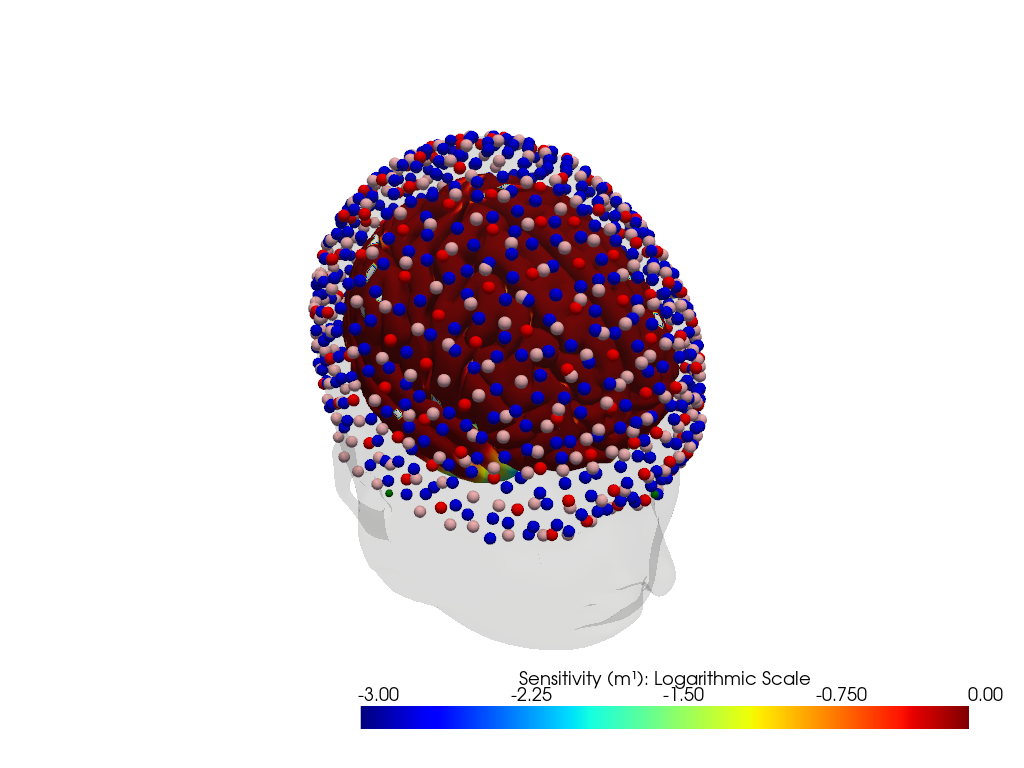
"dataset='ninja_uhd_cap_164x496' headmodel='icbm152'"
Downloading file 'sensitivity_ninja_uhd_cap_164x496_icbm152.nc' from 'https://doc.ibs.tu-berlin.de/cedalion/datasets/dev/sensitivity_ninja_uhd_cap_164x496_icbm152.nc' to '/home/runner/.cache/cedalion/dev'.
References
[7]:
cedalion.bib.dump_to_notebook()
Methods used
| [1] | Fang2009 | cedalion.data.get_precomputed_sensitivity | Qianqian Fang and David A Boas. Monte carlo simulation of photon migration in 3d turbid media accelerated by graphics processing units. Optics express, 17(22):20178–20190, 2009. doi:10.1364/OE.17.020178. |
| [2] | Yu2018 | cedalion.data.get_precomputed_sensitivity | Leiming Yu, Fanny Nina-Paravecino, David Kaeli, and Qianqian Fang. Scalable and massively parallel monte carlo photon transport simulations for heterogeneous computing platforms. Journal of biomedical optics, 23(1):010504–010504, 2018. doi:10.1117/1.JBO.23.1.010504. |
| [3] | Yan2020 | cedalion.data.get_precomputed_sensitivity | Shijie Yan and Qianqian Fang. Hybrid mesh and voxel based monte carlo algorithm for accurate and efficient photon transport modeling in complex bio-tissues. Biomedical Optics Express, 11(11):6262–6270, 2020. doi:10.1364/BOE.409468. |
| [4] | Holmes1998 | cedalion.data.get_colin27_headmodel_files | Colin J. Holmes, Rick Hoge, Louis Collins, Roger Woods, Arthur W. Toga, and Alan C. Evans. Enhancement of mr images using registration for signal averaging. Journal of Computer Assisted Tomography, 22(2):324–333, March 1998. doi:10.1097/00004728-199803000-00032. |
| [5] | Fischl2012 | cedalion.data.get_colin27_headmodel_files, cedalion.data.get_icbm152_headmodel_files | Bruce Fischl. FreeSurfer. NeuroImage, 62(2):774–781, 2012. doi:10.1016/j.neuroimage.2012.01.021. |
| [6] | Schaefer2018 | cedalion.data.get_colin27_headmodel_files, cedalion.data.get_icbm152_headmodel_files | Alexander Schaefer, Ru Kong, Evan M Gordon, Timothy O Laumann, Xi-Nian Zuo, Avram J Holmes, Simon B Eickhoff, and BT Thomas Yeo. Local-global parcellation of the human cerebral cortex from intrinsic functional connectivity mri. Cerebral cortex, 28(9):3095–3114, 2018. doi:10.1093/cercor/bhx179. |
| [7] | Luke2021 | cedalion.data.get_fingertapping | Robert Luke and David McAlpine. fNIRS Finger Tapping Data in BIDS Format. September 2021. doi:10.5281/zenodo.5529797. |
| [8] | Tucker2022 | cedalion.io.snirf.read_snirf | Stephen Tucker, Jay Dubb, Sreekanth Kura, Alexander von Lühmann, Robert Franke, Jörn M. Horschig, Samuel Powell, Robert Oostenveld, Michael Lührs, Édouard Delaire, Zahra M. Aghajan, Hanseok Yun, Meryem A. Yücel, Qianqian Fang, Theodore J. Huppert, Blaise deB. Frederick, Luca Pollonini, David A. Boas, and Robert Luke. Introduction to the shared near infrared spectroscopy format. Neurophotonics, 10(1):013507, 2022. doi:10.1117/1.NPh.10.1.013507. |
| [9] | Fonov2011 | cedalion.data.get_icbm152_headmodel_files | Vladimir Fonov, Alan C. Evans, Kelly Botteron, C. Robert Almli, Robert C. McKinstry, and D. Louis Collins. Unbiased average age-appropriate atlases for pediatric studies. NeuroImage, 54(1):313–327, January 2011. doi:10.1016/j.neuroimage.2010.07.033. |