Constructing 10-10 coordinates on segmented MRI scans
[1]:
# This cells setups the environment when executed in Google Colab.
try:
import google.colab
!curl -s https://raw.githubusercontent.com/ibs-lab/cedalion/dev/scripts/colab_setup.py -o colab_setup.py
# Select branch with --branch "branch name" (default is "dev")
%run colab_setup.py
except ImportError:
pass
[2]:
import cedalion
import cedalion.io
import cedalion.geometry.segmentation
import cedalion.geometry.landmarks
from cedalion.dot import TwoSurfaceHeadModel
import cedalion.data
import os.path
import pyvista
#pyvista.set_jupyter_backend("server")
pyvista.set_jupyter_backend("static")
Load segmentation masks
This example constructs the 10-10 system on the Colin27 average brain.
[3]:
SEG_DATADIR, mask_files, landmarks_file = cedalion.data.get_colin27_segmentation()
masks, t_ijk2ras = cedalion.io.read_segmentation_masks(SEG_DATADIR, mask_files)
/tmp/ipykernel_4941/2857996291.py:1: DeprecationWarning: 'get_colin27_segmentation' is deprecated: This function and the corresponding data files were replaced by cedalion.data.get_icbm152_headmodel_files .
SEG_DATADIR, mask_files, landmarks_file = cedalion.data.get_colin27_segmentation()
Wrap the segmented head with derived surfaces in a TwoSurfaceHeadModel
[4]:
head = TwoSurfaceHeadModel.from_surfaces(
segmentation_dir=SEG_DATADIR,
mask_files = mask_files,
brain_surface_file= os.path.join(SEG_DATADIR, "mask_brain.obj"),
scalp_surface_file= os.path.join(SEG_DATADIR, "mask_scalp.obj"),
landmarks_ras_file=landmarks_file,
brain_face_count=None,
scalp_face_count=None,
smoothing=0.
)
Transform the scalp surface from voxel space (‘ijk’) to RAS space (‘aligned’)
[5]:
scalp_surface = head.scalp
display(scalp_surface)
scalp_surface = scalp_surface.apply_transform(t_ijk2ras)
display(scalp_surface)
TrimeshSurface(faces: 20096 vertices: 10050 crs: ijk units: dimensionless vertex_coords: [])
TrimeshSurface(faces: 20096 vertices: 10050 crs: aligned units: millimeter vertex_coords: [])
Transform initial landmarks from voxel space (‘ijk’) to RAS space (‘aligned’)
[6]:
landmarks_ras = head.landmarks.points.apply_transform(t_ijk2ras)
Construct landmarks
[7]:
lmbuilder = cedalion.geometry.landmarks.LandmarksBuilder1010(scalp_surface, landmarks_ras)
all_landmarks = lmbuilder.build()
/home/runner/work/cedalion/cedalion/src/cedalion/geometry/landmarks.py:319: UserWarning: WIP: distance calculation around ears
warnings.warn("WIP: distance calculation around ears")
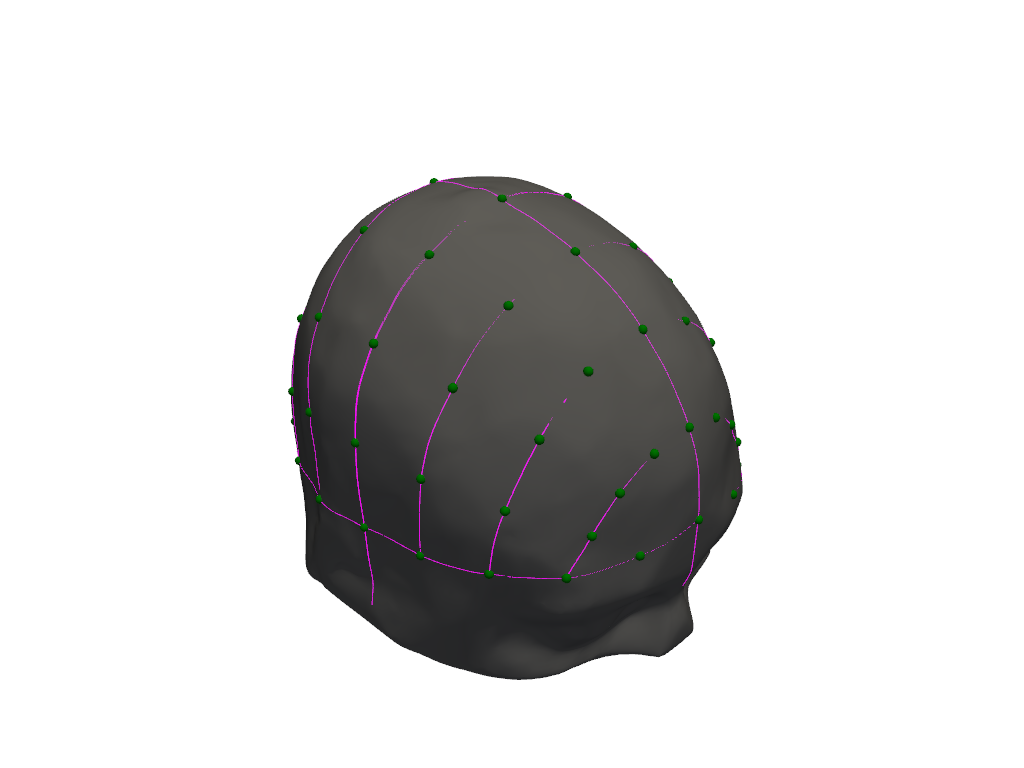
Visualize
[8]:
lmbuilder.plot()
[9]:
display(all_landmarks)
<xarray.DataArray (label: 73, aligned: 3)> Size: 2kB
<Quantity([[ 4.49999997e-03 -1.18565000e+02 -2.30780000e+01]
[-8.60761000e+01 -1.99897000e+01 -4.79860000e+01]
[ 8.57939000e+01 -2.00093000e+01 -4.80310000e+01]
[-6.79195000e-01 8.32593470e+01 -3.94085270e+01]
[ 9.38422944e-01 -9.96505206e+00 9.77001349e+01]
[-8.16800778e+01 -1.74869317e+01 -1.16379427e+01]
[ 8.52482230e+01 -1.73423451e+01 -9.30572974e+00]
[-3.09400847e-01 8.74714115e+01 -1.52475977e+00]
[ 8.56481539e-02 8.06088251e+01 3.60347730e+01]
[ 4.50678552e-01 5.86412576e+01 6.67361432e+01]
[ 7.47354045e-01 2.70835368e+01 8.81769784e+01]
[ 1.04543328e+00 -4.71864907e+01 9.88129122e+01]
[ 9.71563725e-01 -8.12995124e+01 8.27197750e+01]
[ 6.99403531e-01 -1.01130923e+02 5.05509083e+01]
[ 3.71366569e-01 -1.14411044e+02 1.44995041e+01]
[-2.93050952e+01 8.38385564e+01 -7.78311679e+00]
[-5.39362548e+01 6.83548539e+01 -1.21248227e+01]
[-6.84343831e+01 4.20862590e+01 -1.33318018e+01]
[-7.81867340e+01 1.38581944e+01 -1.33136812e+01]
[-8.21228505e+01 -4.58133188e+01 -9.51104301e+00]
...
[ 3.51946921e+01 7.74698875e+01 2.08965702e+01]
[ 4.64388229e+01 7.39021650e+01 5.33104560e+00]
[-7.69866986e+01 -4.64300172e+01 2.86295188e+01]
[-6.37037559e+01 -4.69626082e+01 6.44542386e+01]
[-3.58528885e+01 -4.72507977e+01 9.05794925e+01]
[ 3.89014741e+01 -4.68560358e+01 9.16214907e+01]
[ 6.75609010e+01 -4.62757316e+01 6.66566944e+01]
[ 8.21399827e+01 -4.56004981e+01 3.14858901e+01]
[-6.56979834e+01 -7.60159603e+01 2.66781966e+01]
[-5.24362921e+01 -7.86363208e+01 5.49299789e+01]
[-2.83840012e+01 -8.05154545e+01 7.48536888e+01]
[ 3.25078586e+01 -8.08584983e+01 7.71836848e+01]
[ 5.62515882e+01 -7.90537251e+01 5.69646392e+01]
[ 6.88032125e+01 -7.64611812e+01 2.84188278e+01]
[-4.74219717e+01 -9.87045426e+01 2.01806122e+01]
[-3.55453189e+01 -9.98913547e+01 3.59513955e+01]
[-1.92061655e+01 -1.00841267e+02 4.80375088e+01]
[ 2.00588412e+01 -1.01017798e+02 4.74383218e+01]
[ 3.74483972e+01 -1.00374362e+02 3.70167162e+01]
[ 4.98098774e+01 -9.93390608e+01 2.14747572e+01]], 'millimeter')>
Coordinates:
* label (label) <U3 876B 'Iz' 'LPA' 'RPA' 'Nz' ... 'PO1' 'PO2' 'PO4' 'PO6'
type (label) object 584B PointType.LANDMARK ... PointType.LANDMARK
Dimensions without coordinates: alignedReferences
[10]:
cedalion.bib.dump_to_notebook()
Methods used
| [1] | Holmes1998 | cedalion.data.get_colin27_segmentation | Colin J. Holmes, Rick Hoge, Louis Collins, Roger Woods, Arthur W. Toga, and Alan C. Evans. Enhancement of mr images using registration for signal averaging. Journal of Computer Assisted Tomography, 22(2):324–333, March 1998. doi:10.1097/00004728-199803000-00032. |
| [2] | Oostenveld2001 | cedalion.geometry.landmarks.LandmarksBuilder1010.__init__ | Robert Oostenveld and Peter Praamstra. The five percent electrode system for high-resolution eeg and erp measurements. Clinical Neurophysiology, 112(4):713–719, 2001. doi:https://doi.org/10.1016/S1388-2457(00)00527-7. |